news model
license


























































| ~ | 9203 (C/T) | 9203 (C/A) |
|---|---|---|
| ~ | 9203 (ACA/ATA) | 9203 (ACA/AAA) |
| MitImpact id | MI.1443 | MI.1442 |
| Chr | chrM | chrM |
| Start | 9203 | 9203 |
| Ref | C | C |
| Alt | T | A |
| Gene symbol | MT-ATP6 | MT-ATP6 |
| Extended annotation | mitochondrially encoded ATP synthase membrane subunit 6 | mitochondrially encoded ATP synthase membrane subunit 6 |
| Gene position | 677 | 677 |
| Gene start | 8527 | 8527 |
| Gene end | 9207 | 9207 |
| Gene strand | + | + |
| Codon substitution | ACA/ATA | ACA/AAA |
| AA position | 226 | 226 |
| AA ref | T | T |
| AA alt | M | K |
| Functional effect general | missense | missense |
| Functional effect detailed | missense | missense |
| OMIM id | 516060 | 516060 |
| HGVS | NC_012920.1:g.9203C>T | NC_012920.1:g.9203C>A |
| HGNC id | 7414 | 7414 |
| Respiratory Chain complex | V | V |
| Ensembl gene id | ENSG00000198899 | ENSG00000198899 |
| Ensembl transcript id | ENST00000361899 | ENST00000361899 |
| Ensembl protein id | ENSP00000354632 | ENSP00000354632 |
| Uniprot id | P00846 | P00846 |
| Uniprot name | ATP6_HUMAN | ATP6_HUMAN |
| Ncbi gene id | 4508 | 4508 |
| Ncbi protein id | YP_003024031.1 | YP_003024031.1 |
| PhyloP 100V | 1.566 | 1.566 |
| PhyloP 470Way | 0.724 | 0.724 |
| PhastCons 100V | 0.003 | 0.003 |
| PhastCons 470Way | 0.556 | 0.556 |
| PolyPhen2 | possibly_damaging | benign |
| PolyPhen2 score | 0.78 | 0.14 |
| SIFT | deleterious | deleterious |
| SIFT score | 0 | 0.01 |
| SIFT4G | Damaging | Damaging |
| SIFT4G score | 0.0 | 0.0 |
| VEST | Neutral | Neutral |
| VEST pvalue | 0.38 | 0.37 |
| VEST FDR | 0.65 | 0.65 |
| Mitoclass.1 | damaging | damaging |
| SNPDryad | Pathogenic | Neutral |
| SNPDryad score | 0.93 | 0.87 |
| MutationTaster | . | Polymorphism |
| MutationTaster score | . | 1.0 |
| MutationTaster converted rankscore | . | 0.08975 |
| MutationTaster model | . | simple_aae |
| MutationTaster AAE | . | T226K |
| fathmm | . | Tolerated |
| fathmm score | . | 4.63 |
| fathmm converted rankscore | . | 0.01817 |
| AlphaMissense | ambiguous | likely_pathogenic |
| AlphaMissense score | 0.3999 | 0.6629 |
| CADD | Deleterious | Deleterious |
| CADD score | 4.026788 | 2.829472 |
| CADD phred | 23.6 | 21.5 |
| PROVEAN | Damaging | Damaging |
| PROVEAN score | -3.41 | -3.85 |
| MutationAssessor | . | medium |
| MutationAssessor score | . | 3.145 |
| EFIN SP | Neutral | Neutral |
| EFIN SP score | 0.888 | 0.884 |
| EFIN HD | Damaging | Damaging |
| EFIN HD score | 0.258 | 0.262 |
| MLC | Neutral | Neutral |
| MLC score | 0.10477398 | 0.10477398 |
| PANTHER score | . | . |
| PhD-SNP score | . | . |
| APOGEE1 | Neutral | Neutral |
| APOGEE1 score | 0.44 | 0.39 |
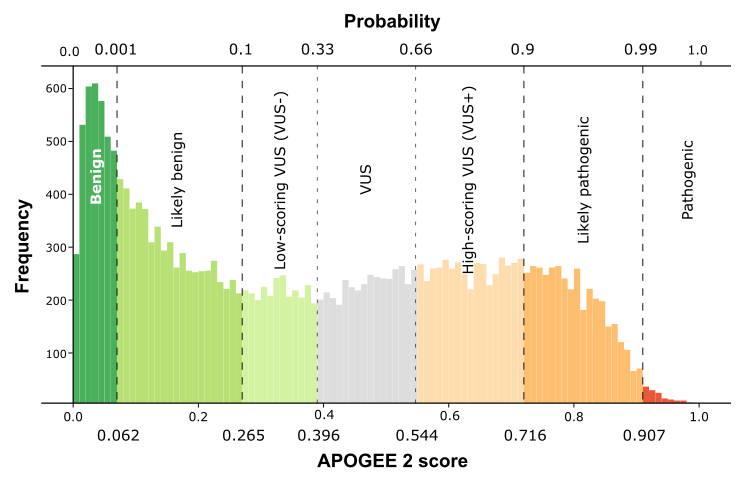
| APOGEE2 | Likely-benign | Likely-benign |
| APOGEE2 score | 0.224752409599884 | 0.203603946794222 |
| CAROL | deleterious | deleterious |
| CAROL score | 1 | 0.99 |
| Condel | neutral | neutral |
| Condel score | 0.11 | 0.44 |
| COVEC WMV | deleterious | deleterious |
| COVEC WMV score | 4 | 1 |
| MtoolBox | deleterious | neutral |
| MtoolBox DS | 0.77 | 0.33 |
| DEOGEN2 | . | Tolerated |
| DEOGEN2 score | . | 0.083704 |
| DEOGEN2 converted rankscore | . | 0.37114 |
| Meta-SNP | . | . |
| Meta-SNP score | . | . |
| PolyPhen2 transf | low impact | medium impact |
| PolyPhen2 transf score | -1.28 | -0.01 |
| SIFT_transf | low impact | medium impact |
| SIFT transf score | -1.4 | -0.84 |
| MutationAssessor transf | medium impact | medium impact |
| MutationAssessor transf score | 0.85 | 1.15 |
| CHASM | Neutral | Neutral |
| CHASM pvalue | 0.76 | 0.63 |
| CHASM FDR | 0.9 | 0.9 |
| ClinVar id | . | . |
| ClinVar Allele id | . | . |
| ClinVar CLNDISDB | . | . |
| ClinVar CLNDN | . | . |
| ClinVar CLNSIG | . | . |
| MITOMAP Disease Clinical info | . | . |
| MITOMAP Disease Status | . | . |
| MITOMAP Disease Hom/Het | ./. | ./. |
| MITOMAP General GenBank Freq | 0.0% | . |
| MITOMAP General GenBank Seqs | 0 | . |
| MITOMAP General Curated refs | . | . |
| MITOMAP Variant Class | polymorphism | . |
| gnomAD 3.1 AN | 56434.0 | . |
| gnomAD 3.1 AC Homo | 0.0 | . |
| gnomAD 3.1 AF Hom | 0.0 | . |
| gnomAD 3.1 AC Het | 2.0 | . |
| gnomAD 3.1 AF Het | 3.54396e-05 | . |
| gnomAD 3.1 filter | PASS | . |
| HelixMTdb AC Hom | 1.0 | . |
| HelixMTdb AF Hom | 5.1024836e-06 | . |
| HelixMTdb AC Het | 0.0 | . |
| HelixMTdb AF Het | 0.0 | . |
| HelixMTdb mean ARF | . | . |
| HelixMTdb max ARF | . | . |
| ToMMo 54KJPN AC | 1 | . |
| ToMMo 54KJPN AF | 1.8e-05 | . |
| ToMMo 54KJPN AN | 54302 | . |
| COSMIC 90 | . | . |
| dbSNP 156 id | . | . |